
|
GENTAUR EUROPE BELGIUM1 tel +32 2 732 5688 fax +32 2 732 4414 [email protected] Av. de l' Armée 68 B-1040 Brussels France tel 01 43 25 01 50 fax 01 43 25 01 60 9, rue Lagrange 75005 Paris Italy tel 02 36 00 65 93 fax +32 16 50 90 45 20135 Milano Germany tel +32 16 58 90 45 fax +32 16 50 90 45 Forckenbeckstraße 6 D-52074 Aachen Japan tel +81 78 386 0860 fax +81 78 306 0296 Minaatojimaminami-manchi Chuo-ku, Kobe 065-0047 |
| Protein Identification |
|
You provide us with a,b, or c:
Required Information:
Please ship the protein without cooling (dried gel bands or spots, lyophilized protein) or on blue ice (ammonium sulfate precipitate) to:
|
|
Our service:
|
|
Order Information:
Please note: In case the protein amount or purity was not sufficient to perform MALDI-TOF MS, a set up fee of € 175 is charged. |
| Examples: |
| The SDS-PAGE gel illustration below shows the results of a protein purification (upper band) by metal-chelate chromatography after trying different washing conditions applied to eliminate the lower band (impurity). |
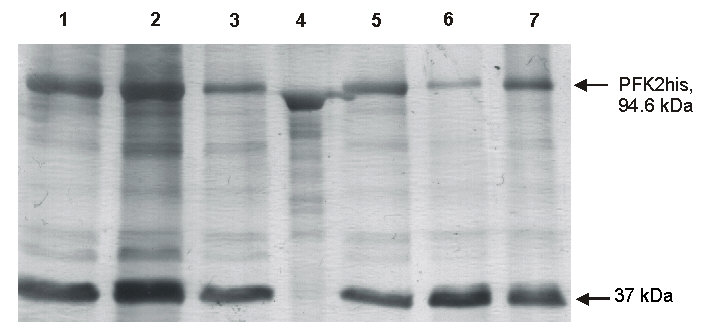
|
| The MALDI-TOF MS spectrum of the protein preparation confirmed the results obtained with the SDS-PAGE. |
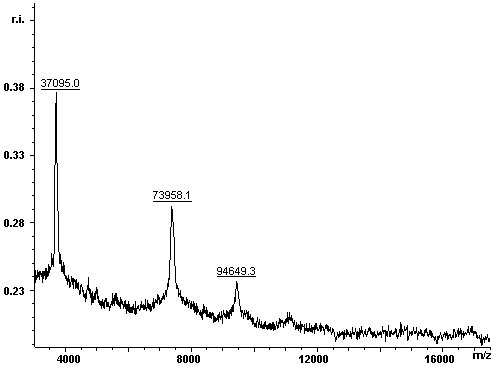
|
|
The contaminant has a molecular weight of 37095 Da. To efficiently eliminate the impurity, the 37 kDa protein was identified by MALDI-TOF MS of a tryptic protein digest: The lower band was excised from the gel and the in gel-digestion was carried out with trypsin. The tryptic digest was analysed with MALDI-TOF MS. The resulting mass spectra are given below: |
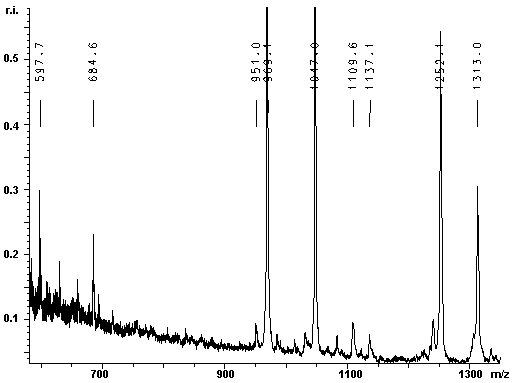
|
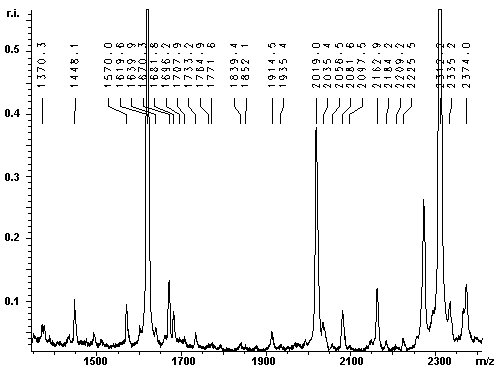
|
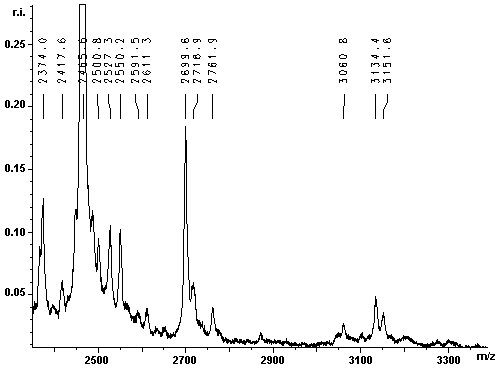
|
| Protein Candidates | |||
|
Rank
|
Probability
|
Protein Description |
MW (kDa)
|
|
1
|
1.0e+00
|
gi|6324486|ref|NP_014555.1| Alcohol dehydrogenase; Adh1p [Saccharomyces cerevisiae] |
37
|
|
2
|
3.2e-03
|
gi|1168350|sp|P00330|ADH1_YEAST ALCOHOL DEHYDROGENASE I |
37
|
|
3
|
7.8e-09
|
gi|6324174|ref|NP_014244.1| Ynl155wp [Saccharomyces cerevisiae] |
32
|
|
4
|
1.1e-09
|
gi|7245674|pdb|1YLV|A Chain A, Schiff-Base Complex Of Yeast 5-Aminolaevulinic Acid Dehydratase With Laevulinic Acid |
38
|
|
5
|
3.9e-10
|
gi|171974|gb|AAA34790.1| (M81696) mitochondrial ribosomal protein [Saccharomyces cerevisiae] |
29
|
|
6
|
3.2e-10
|
gi|6325350|ref|NP_015418.1| Ypr093cp [Saccharomyces cerevisiae] |
36
|
|
7
|
3.0e-10
|
gi|223142|prf||0512226A dehydrogenase isozyme I,alcohol [Saccharomyces cerevisiae] |
32
|
|
8
|
3.0e-10
|
gi|6320613|ref|NP_010693.1| 263-amino acid mitochondrial ribosomal large subunit protein; similar to L23 family of ribosomal proteins; Mrp20p [Saccharomyces cerevisiae] |
31
|
|
9
|
1.2e-10
|
gi|626218|pir||S45958 probable membrane protein YBR090c - yeast (Saccharomyces cerevisiae) |
13
|
|
10
|
1.2e-10
|
gi|6319560|ref|NP_009642.1| mitochondrial ADP/ATP translocator; Aac3p [Saccharomyces cerevisiae] |
33
|
| The contaminating protein was identified as alcohol dehydrogenase and could now be removed easily from the mixture by exploiting its known hydrophobicity (washing the column with isopropanol). | |||
Our service:
|
|
Order Information:
|

